|
Getting your Trinity Audio player ready…
|
We have already looked at three powerful biological parallels:
Endosymbiosis: independent systems become internal modules, like mitochondria inside cells or tools/plugins/retrievers inside AI systems.
Horizontal gene transfer: useful information jumps across lineages, like models borrowing capabilities through fine-tuning, distillation, synthetic data, tool use, or shared embeddings.
Epigenetics: the genome stays mostly fixed, but expression changes with context, like frozen LLM weights producing different behavior through prompts, context windows, system instructions, retrieval, and activation patterns.
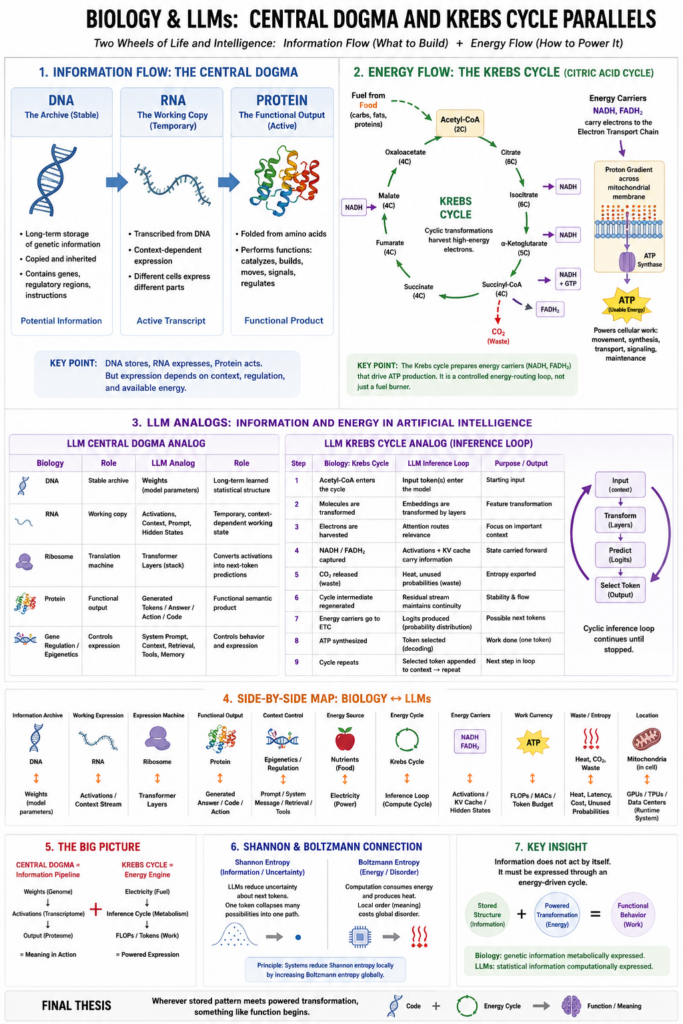
Now we come to two even deeper biological structures:
The central dogma of biology:
DNA → RNA → Protein
And the Krebs cycle, also called the citric acid cycle:
nutrients → acetyl-CoA → cyclic chemical transformations → energy carriers → ATP production
These are different kinds of systems.
The central dogma is about information flow.
The Krebs cycle is about energy flow.
Together, they form one of the deepest truths in biology:
Life requires both a code that can be interpreted and an energy cycle that pays for the interpretation.
That is exactly where the LLM analogy becomes interesting.
An LLM is not alive. It does not metabolize. It does not have DNA. It does not reproduce biologically. But it does have something structurally similar:
Stored information, contextual transcription, active expression, and energy-costly computation.
In plain English:
Biology turns genetic code into living action.
LLMs turn stored statistical structure into contextual meaning.
I. The Central Dogma in Biology
The central dogma says that biological information usually flows like this:
DNA → RNA → Protein
DNA is the long-term archive.
RNA is the working copy.
Protein is the functional output.
That is the simplest version. Of course, real biology is full of complications: regulatory RNAs, reverse transcription, epigenetics, alternative splicing, RNA editing, feedback loops, protein modifications, and environmental effects. But the core picture is still powerful.
DNA: the stable archive
DNA stores information across generations.
It is relatively stable.
It is protected.
It is copied.
It contains genes, regulatory regions, instructions, switches, and patterns.
But DNA does not usually “do” the work directly. DNA is more like a library, blueprint, recipe collection, or compressed archive.
A gene sitting in DNA is potential information.
It becomes biologically meaningful only when expressed.
That matters for our LLM analogy.
An LLM’s trained weights are like DNA in one important sense:
The weights are stored learning.
They are not a database of explicit facts in the ordinary sense. They are not a library of sentences. They are a vast compressed statistical geometry formed by training.
The model’s weights are the long-term encoded result of exposure to data.
So in our analogy:
DNA ≈ trained model weights
Not because weights are literally genes, but because both are durable encoded structures that can generate behavior when activated.
II. RNA as the Working Copy
In biology, DNA is transcribed into RNA.
RNA is more temporary, more flexible, more situational.
A cell does not express all genes all the time. It selectively transcribes certain genes into RNA depending on context.
A liver cell, a neuron, and a muscle cell contain essentially the same DNA, but they express different parts of it.
That is the crucial point.
The genome is mostly fixed, but expression is conditional.
This is very close to how an LLM behaves.
The LLM has fixed weights during inference, but a prompt activates only certain pathways, associations, concepts, and capabilities.
The prompt is like a cellular condition that says:
“Express this part of the stored structure now.”
In an LLM, the working equivalent of RNA is not one single thing. It is more like a combination of:
the prompt, the context window, the token stream, the hidden states, the attention patterns, and the temporary activations.
These are transient. They exist during the “living moment” of inference.
They are not the model’s permanent memory.
They are the model’s active expression.
So:
RNA ≈ context-dependent activation stream
Or more specifically:
RNA ≈ the temporary working transcript produced when fixed weights meet a prompt.
The model’s weights contain learned potential.
The prompt causes part of that potential to be transcribed into active computation.
III. Protein as Functional Output
Proteins are where much of biological function happens.
They build structures.
They catalyze reactions.
They move molecules.
They form channels.
They receive signals.
They regulate other systems.
They are the “doing” molecules of the cell.
DNA stores.
RNA carries.
Protein acts.
In LLMs, the closest analog to protein is the output: the generated text, code, plan, explanation, image prompt, tool call, or action instruction.
The output is where the model’s internal structure becomes externally functional.
So:
Protein ≈ generated output or executable action
But we can go one level deeper.
A protein is not merely an output string. It folds into a functional shape. Its sequence matters because the sequence determines the structure, and the structure determines the function.
That is strongly analogous to language.
An LLM generates token sequences. But the value is not merely in the tokens individually. The value is in the formed structure:
a sentence, an explanation, a program, a proof, a plan, a metaphor, a diagnosis, a design.
In biology:
amino acid sequence → folded protein → biological function
In LLMs:
token sequence → semantic structure → cognitive function
This is a very strong analogy.
A protein is a folded molecular tool.
A good LLM answer is a folded semantic tool.
IV. The Central Dogma as an LLM Pipeline
Here is the biological version:
DNA → RNA → Protein
Here is an LLM-like version:
Weights → Activations → Output
Or more fully:
Training data compressed into weights → prompt-conditioned hidden states → generated tokens/actions
That gives us this map:
| Biology | Role | LLM Analog | Role |
|---|---|---|---|
| DNA | Long-term encoded information | Weights | Long-term learned structure |
| Gene | Expressible information unit | Learned feature/capability | Expressible model pattern |
| RNA | Temporary working copy | Prompt/context/activations | Temporary inference state |
| Ribosome | Translation machine | Transformer layers | Convert active state into output |
| Protein | Functional product | Generated answer/action/code | Functional semantic product |
| Gene regulation | Controls expression | System prompt/context/retrieval | Controls behavior |
| Epigenetics | Context-sensitive expression | Prompting, instruction hierarchy, memory, retrieval | Context-sensitive model expression |
The deepest summary is this:
In biology, DNA does not become life until it is expressed.
In LLMs, weights do not become thought-like behavior until they are activated.
That connects beautifully to your earlier phrase:
Weights are frozen learning; activations are living thought.
The central dogma version would be:
Weights are the genome; activations are the transcriptome; outputs are the proteome of meaning.
V. But There Is a Major Difference
We must be careful.
In biology, DNA is a symbolic molecular code. Codons map to amino acids through cellular machinery.
In LLMs, weights are not a clean symbolic code. They are distributed numerical relationships.
A gene can often be identified as a physical sequence.
A model capability is usually spread across many weights, layers, attention heads, MLP neurons, and activation subspaces.
So the analogy is not:
“A model has genes exactly like DNA.”
The better analogy is:
“A model has stored generative potential, and inference is context-dependent expression of that potential.”
Biology is chemically embodied.
LLMs are computationally embodied.
Biology uses carbon chemistry.
LLMs use matrix multiplication.
Biology pays energy in ATP.
LLMs pay energy in electricity.
But both systems depend on this same deep architecture:
stable stored structure + context-sensitive expression + energy-costly transformation = functional output
That brings us to the Krebs cycle.
VI. The Krebs Cycle: Biology’s Energy Wheel
The central dogma explains how biological information becomes function.
The Krebs cycle explains how biological energy is harvested to pay for function.
The Krebs cycle is a circular pathway inside mitochondria. Its job is to take carbon-based fuel and strip out high-energy electrons.
Those electrons are transferred to carrier molecules, mainly NADH and FADH₂.
Those carriers then feed the electron transport chain, which helps create a proton gradient across the mitochondrial membrane. That proton gradient drives ATP synthase, producing ATP.
In plain English:
The Krebs cycle does not directly make most of the ATP.
It prepares charged energy carriers that feed the larger ATP-producing system.
This matters.
The Krebs cycle is not merely a “fuel burner.”
It is a regulated energy-routing system.
It turns food-derived molecules into reusable energy currency.
A simplified version:
glucose/fat/protein → acetyl-CoA → Krebs cycle → NADH/FADH₂ → electron transport chain → proton gradient → ATP
So where is the LLM analogy?
VII. The LLM Version of the Krebs Cycle
An LLM also requires energy to turn stored structure into output.
But the energy does not come from glucose.
It comes from electricity.
The “metabolic machinery” is the hardware:
GPUs, TPUs, NPUs, memory systems, data centers, cooling systems, power grids, and increasingly edge devices.
The LLM’s Krebs-like cycle is not chemical. It is computational.
At inference time, the system repeatedly performs something like this:
token input → embedding → layer transformations → attention/MLP operations → logits → next token → new input → repeat
That is cyclic.
Each generated token becomes part of the context for the next token.
So the LLM has a loop:
token → hidden state → probability distribution → selected token → updated context → token
This is not the Krebs cycle chemically, but it is similar in architecture:
a cyclic transformation process that converts input into usable work by repeatedly passing through structured stages.
In biology:
acetyl-CoA enters the Krebs cycle, and energy carriers emerge.
In LLM inference:
tokens enter the transformer stack, and probability-shaped semantic outputs emerge.
The energy carriers in biology are NADH and FADH₂.
The “energy carriers” in LLMs are more abstract:
activations, attention scores, residual streams, logits, KV cache entries, and updated context.
They carry forward the usable computational state.
VIII. Two Wheels: One for Meaning, One for Energy
Now we can place the central dogma and Krebs cycle side by side.
Biology has:
DNA → RNA → Protein
nutrient fuel → Krebs cycle → ATP
LLMs have:
weights → activations → output
electricity → matrix operations → inference cycles
The central dogma is the information-expression pipeline.
The Krebs cycle is the energy-conversion loop.
Together:
| Biology | LLM Analog |
|---|---|
| DNA stores biological information | Weights store learned statistical structure |
| RNA expresses selected information | Prompt/context activates selected model pathways |
| Protein performs biological work | Output performs semantic/cognitive work |
| Nutrients provide fuel | Electricity provides fuel |
| Krebs cycle routes energy | Hardware computation routes energy |
| NADH/FADH₂ carry electrons | Activations/KV cache carry computational state |
| ATP powers cellular work | FLOPs/MACs power inference work |
| Mitochondria supply energy | GPUs/NPUs/data centers supply compute |
The big idea:
The central dogma tells life what to build.
The Krebs cycle helps pay the energy cost of building and acting.
For LLMs:
The weights tell the model what kinds of meaning can be expressed.
The compute cycle pays the energy cost of expressing that meaning.
IX. The Ribosome and the Transformer
The ribosome deserves special attention.
In biology, the ribosome translates RNA into protein.
It reads an RNA sequence and builds an amino acid chain.
That chain folds into a protein.
The ribosome is not DNA. It is not RNA. It is not protein output. It is the machine that converts one representational form into another.
That sounds very much like the transformer stack.
The transformer converts token embeddings and contextual activations into next-token predictions.
In biology:
RNA sequence enters ribosome → amino acid chain emerges
In LLMs:
token/context sequence enters transformer → probability distribution over next tokens emerges
The ribosome translates from nucleotide language to amino acid language.
The transformer translates from token-context geometry to semantic continuation.
Again, not literally the same, but structurally similar.
The ribosome is a biological translator.
The transformer is a mathematical translator.
The ribosome asks:
Given this RNA sequence, what protein chain should be assembled?
The LLM asks:
Given this token context, what next token best continues the pattern?
But the larger meaning is:
Both systems use a stable interpreter to convert encoded information into functional output.
X. The Central Dogma Is Not a One-Way Dictatorship
The phrase “central dogma” can sound rigid: DNA makes RNA makes protein, end of story.
But biology is more interactive than that.
Proteins regulate DNA.
RNA can regulate other RNA.
The environment affects gene expression.
Chemical marks affect transcription.
Signals from metabolism influence what genes are expressed.
Stress, nutrients, hormones, temperature, toxins, and developmental stage can all alter expression.
So the real picture is not a simple straight line.
It is more like:
stored code → working expression → functional product → feedback into regulation
That is also true for LLM systems.
A simple LLM inference pass looks one-way:
prompt → output
But modern AI systems increasingly include feedback:
prompt → output → tool use → retrieved data → revised context → new output
Or:
output → user correction → new prompt → different activation pattern
Or:
model output → synthetic training data → future model update
So the central dogma analogy becomes even richer when we include feedback.
The model’s weights are fixed during one inference run, but the broader AI ecosystem is not fixed.
There is:
fine-tuning, reinforcement learning, retrieval augmentation, memory, tool use, evaluation, distillation, and model updating.
That resembles the biological reality that gene expression is embedded in regulatory loops.
XI. Krebs Cycle as a Semantic Energy Ratchet
This is where your entropy framework becomes especially powerful.
The Krebs cycle is not just a wheel. It is a ratchet.
It takes a high-energy molecule and gradually extracts useful energy in controlled steps.
It does not explode the fuel all at once.
It does not waste the gradient.
It captures the gradient.
That is life’s genius.
Life does not merely burn.
Life meters energy through structure.
That is exactly what a cell does:
It prevents energy from becoming mere heat too quickly.
It channels energy through molecular machinery.
It extracts usable work before entropy wins.
In your language:
Life rides Boltzmann entropy while reducing Shannon entropy locally.
The Krebs cycle participates in that bargain.
It takes energetic disorder-potential and turns it into organized cellular work.
Now compare that with LLM inference.
An LLM does not merely emit random tokens.
It uses energy-consuming computation to reduce uncertainty over the next token.
At each step, the model begins with a vast possible space of continuations.
Then the network narrows the space.
It creates a probability distribution.
The selected token reduces uncertainty.
Then the next step begins.
That is a Shannon entropy reduction process.
But it costs Boltzmann energy: electricity, heat, cooling, hardware wear, power-grid load.
So we can say:
The LLM’s inference loop is a semantic entropy ratchet.
It uses physical energy to reduce uncertainty in symbolic space.
Biology:
food gradient → chemical cycle → ATP → organized life
LLM:
electrical gradient → compute cycle → token selection → organized meaning
This is not metaphor only. It is structurally serious.
Both systems require physical energy to produce ordered information-processing behavior.
XII. The Krebs Cycle and the Transformer Loop
Let’s compare them as cycles.
Krebs cycle
- Acetyl-CoA enters.
- Carbon molecules are rearranged.
- Electrons are harvested.
- NADH and FADH₂ carry energy onward.
- CO₂ is released.
- The cycle regenerates its starting molecule.
- The process repeats.
Transformer inference loop
- A token/context enters.
- Embeddings are transformed.
- Attention redistributes relevance.
- MLPs expand and compress features.
- Logits are produced.
- A token is selected.
- The token is appended to context.
- The process repeats.
The structural analogy:
| Krebs Cycle | Transformer Inference |
|---|---|
| Acetyl-CoA enters cycle | Token/context enters model |
| Molecular intermediates transform | Hidden states transform |
| Electrons are extracted | Predictive signal is extracted |
| NADH/FADH₂ carry energy forward | KV cache/activations carry context forward |
| CO₂ is waste output | Heat/unused probabilities are waste |
| Oxaloacetate regenerated | Context loop continues |
| Cycle repeats | Token generation repeats |
The Krebs cycle is a chemical recurrence.
LLM inference is a computational recurrence.
Both are structured loops that prevent raw input from dissipating uselessly.
XIII. Protein Folding and Meaning Folding
There is another beautiful bridge.
The central dogma does not end at protein sequence.
A chain of amino acids must fold into a working shape.
The sequence is one-dimensional.
The folded protein is three-dimensional.
The function depends on the fold.
Similarly, an LLM output is a one-dimensional sequence of tokens, but the meaning is not one-dimensional.
The meaning “folds” into a conceptual structure in the reader’s mind.
A good explanation is like a protein.
Its words are the sequence.
Its concept is the fold.
Its usefulness is the function.
This is why bad writing can contain correct words but fail as thought.
It has sequence without functional folding.
A protein with the wrong fold may be useless or harmful.
An answer with the wrong conceptual fold may be confusing or misleading.
So:
DNA encodes protein sequence, but life depends on folded function.
LLMs generate token sequence, but intelligence depends on folded meaning.
This is one of the strongest analogies yet.
XIV. The Central Dogma of LLMs
If we were to write an equivalent “central dogma” for LLMs, it might be:
Training data shapes weights.
Prompts activate weights.
Activations generate tokens.
Tokens fold into meaning.
Or more compactly:
Data → Weights → Activations → Tokens → Meaning
But there is an important distinction.
In biology, the central dogma starts with DNA because DNA is the inherited archive.
In LLMs, the original “ancestral environment” is training data.
So the full LLM version is:
Corpus → Weights → Contextual Activations → Output
The corpus is like evolutionary history.
The weights are like the genome.
The prompt is like the cellular environment.
The activations are like RNA expression.
The output is like protein function.
The user response is like environmental feedback.
A refined LLM central dogma:
Learned statistical inheritance becomes contextual expression, which becomes functional semantic behavior.
Or in your preferred style:
The LLM genome is not made of DNA.
It is made of weights.
Its transcriptome is the activation stream.
Its proteome is the generated world of language, code, plans, and symbols.
XV. The Krebs Cycle of LLMs
Now let’s write an equivalent “Krebs cycle” for LLMs.
Biology:
Nutrient energy enters as acetyl-CoA and is transformed into electron carriers that power ATP generation.
LLM:
Electrical energy enters hardware and is transformed through matrix operations into activation states that power token generation.
A possible LLM Krebs cycle:
- Electricity powers hardware.
- Hardware performs matrix multiplications.
- Matrix multiplications transform token embeddings.
- Attention routes contextual relevance.
- MLPs reshape feature space.
- Residual streams preserve and combine information.
- Logits form a probability landscape.
- A token is selected.
- The selected token re-enters the loop as new context.
That is the semantic metabolism of inference.
It is not life, but it is metabolism-like in structure:
input, transformation, energy cost, intermediate carriers, output, recurrence.
The central dogma of LLMs explains what is expressed.
The Krebs-like inference cycle explains how expression is powered.
XVI. Central Dogma + Krebs Cycle = Information Needs Energy
This is the deepest bridge.
Biology discovered that information is not enough.
DNA by itself is inert.
A dead cell may still contain DNA.
The DNA may still be readable.
But without metabolism, membrane gradients, ribosomes, enzymes, ATP, and regulation, the information does not live.
Likewise, model weights sitting on a disk are inert.
They may contain astonishing learned structure.
But without electricity, hardware, memory bandwidth, inference code, and input context, they do nothing.
So both biology and AI teach the same principle:
Information requires an energetic interpreter.
DNA needs the cell.
Weights need the machine.
Genes need transcription.
Weights need activation.
RNA needs ribosomes.
Activations need transformer layers.
Proteins need ATP-driven cellular context.
Outputs need computational infrastructure and user interpretation.
This is why the central dogma and Krebs cycle belong together.
One is the code.
The other is the engine.
One is Shannon.
The other is Boltzmann.
XVII. Shannon and Boltzmann in This Picture
Your larger theory fits here naturally.
Shannon entropy
Shannon entropy concerns uncertainty, information, signal, and possible messages.
In biology, DNA reduces uncertainty by storing instructions. Gene expression selectively narrows what the cell does.
In LLMs, token prediction reduces uncertainty over possible continuations.
Each generated token collapses a cloud of possible meanings into an actual path.
Boltzmann entropy
Boltzmann entropy concerns physical disorder, heat, molecular states, and thermodynamic dispersal.
In biology, metabolism exports heat and waste while maintaining local organization.
In LLMs, computation consumes electricity and emits heat while producing ordered symbolic output.
So the central pattern is:
A system reduces Shannon entropy locally by increasing Boltzmann entropy globally.
A cell reduces uncertainty about biological action by burning fuel.
An LLM reduces uncertainty about symbolic continuation by burning electricity.
That is the bridge.
The central dogma is the Shannon side.
The Krebs cycle is the Boltzmann side.
Together they say:
Meaning is not free.
Order is not free.
Interpretation is not free.
Life and intelligence both pay thermodynamic rent.
XVIII. Why the Krebs Cycle Is More Than “Fuel”
The Krebs cycle also produces building-block precursors.
It is amphibolic: both catabolic and anabolic.
That means it breaks things down for energy and also supplies ingredients for biosynthesis.
This is important for the LLM analogy.
Inference is not merely output generation. In modern systems, inference can also produce material for future learning:
synthetic data, summaries, plans, code, annotations, labels, evaluations, embeddings, and training examples.
So the AI compute cycle can be both:
catabolic: consuming energy and input to produce output
and
anabolic: creating new informational structures that can be reused later
Biology uses metabolism to maintain and rebuild itself.
AI systems increasingly use inference to generate data that improves future systems.
That resembles a primitive informational metabolism.
Again, not alive, but increasingly life-like in architecture.
XIX. A Powerful Composite Map
Here is the integrated map:
| Biological System | Function | LLM/AI Analog | Function |
|---|---|---|---|
| Evolution | Long-term search over forms | Training | Long-term search over parameters |
| DNA | Stable inherited code | Weights | Stable learned structure |
| Gene | Expressible functional unit | Feature/capability | Expressible learned pattern |
| Epigenetics | Context controls expression | Prompt/system/context | Context controls behavior |
| RNA | Temporary transcript | Activations/context stream | Temporary working state |
| Ribosome | Translation engine | Transformer stack | Token-generation engine |
| Protein | Functional output | Answer/code/action | Functional semantic object |
| Protein folding | Sequence becomes structure | Meaning formation | Tokens become concept |
| Nutrients | Energy source | Electricity | Energy source |
| Krebs cycle | Energy-routing loop | Inference loop | Compute-routing loop |
| NADH/FADH₂ | Energy carriers | Activations/KV cache | State carriers |
| ATP | Work currency | FLOPs/MACs/token budget | Compute currency |
| Heat/waste | Exported entropy | Heat/latency/cost | Exported entropy |
| Cell | Embodied interpreter | Runtime system | Embodied computational interpreter |
This is the cleanest combined statement:
Biology stores information in DNA, expresses it through RNA, realizes it in protein, and powers the whole process through metabolism.
LLMs store learned structure in weights, express it through activations, realize it in generated outputs, and power the whole process through computation.
XX. The Big Insight
The central dogma and the Krebs cycle are usually taught separately.
One belongs to genetics.
The other belongs to metabolism.
But life does not separate them.
A cell is not just information.
A cell is not just energy.
A cell is information riding energy.
That is also the key to understanding AI systems.
An LLM is not just a pile of weights.
It is not just electricity.
It is stored statistical structure riding an energy-consuming inference process.
So the biological parallel is:
DNA without metabolism is a frozen archive.
Weights without compute are frozen learning.
And:
Metabolism without information is mere chemistry.
Compute without learned structure is mere heat.
Life emerges when information and energy are coupled.
AI behavior emerges when weights and compute are coupled.
This is why the analogy is so striking.
XXI. The Central Dogma as “Semantic Genetics”
Let us name the LLM version:
Semantic genetics
In biology:
DNA carries genetic possibility.
In LLMs:
weights carry semantic possibility.
In biology:
expression depends on cell type, environment, signals, and regulation.
In LLMs:
expression depends on prompt, system message, context window, retrieval, tools, and user intent.
In biology:
protein function depends on folding.
In LLMs:
answer function depends on conceptual coherence.
So we might say:
The LLM does not remember in sentences.
It inherits a semantic genome.
The prompt transcribes a temporary working state.
The transformer translates that state into token sequences.
The reader folds those tokens into meaning.
That is a beautiful central dogma of machine cognition.
XXII. The Krebs Cycle as “Semantic Metabolism”
Now name the LLM version of the Krebs cycle:
Semantic metabolism
In biology, metabolism is not just energy use. It is controlled transformation.
In LLMs, inference is controlled transformation.
A prompt does not simply pass through the model unchanged. It is embedded, transformed, attended to, normalized, expanded, compressed, and projected into probabilities.
This is very much like metabolism:
Molecules enter, are transformed through staged pathways, and emerge as usable work or building blocks.
For LLMs:
Tokens enter, are transformed through staged layers, and emerge as usable meaning or action.
The transformer block is a metabolic chamber of meaning.
Attention routes relevance.
The MLP transforms features.
The residual stream preserves continuity.
LayerNorm stabilizes flow.
The KV cache carries forward context.
The logits expose possible futures.
Sampling or decoding selects one path.
Then the selected token re-enters the cycle.
That is semantic metabolism.
XXIII. Where the Analogy Breaks
We should be precise about the limits.
Biology is self-maintaining. LLMs are not.
Biology builds its own parts. LLMs do not autonomously manufacture GPUs.
Biology reproduces. LLMs do not biologically reproduce.
Biology has membranes. LLMs have system boundaries, but not living membranes.
Biology has homeostasis. LLMs have runtime stability, but not organismic homeostasis.
Biology has metabolism in the literal chemical sense. LLMs consume energy indirectly through hardware.
So the analogy should not become mystical.
The correct claim is not:
“LLMs are alive.”
The better claim is:
“LLMs show an abstract version of the same architecture: stored information, context-sensitive expression, energy-costly transformation, and functional output.”
That is already profound enough.
XXIV. The Most Striking Parallel
The strongest parallel may be this:
DNA does not directly specify the living organism.
It specifies a generative process.
Likewise:
LLM weights do not directly store all possible answers.
They specify a generative process.
DNA is not a complete list of body parts.
Weights are not a complete list of sentences.
DNA participates in a developmental system.
Weights participate in an inference system.
The organism emerges through expression, regulation, environment, and energy flow.
The answer emerges through prompting, activation, decoding, context, and compute.
That means both biology and LLMs are not lookup tables.
They are generative engines.
XXV. Final Condensed Thesis
The central dogma gives us the information pipeline:
DNA → RNA → Protein
The Krebs cycle gives us the energy engine:
Fuel → cyclic transformation → usable work
The LLM analog is:
Weights → activations → output
and:
Electricity → matrix computation → token generation
Together:
Stored structure + contextual expression + energy flow = functional behavior
For biology, that behavior is living action.
For LLMs, that behavior is semantic action.
So the clean final formulation is:
Biology is genetic information metabolically expressed.
LLMs are statistical information computationally expressed.
Or, in your entropy language:
Life uses metabolism to spend Boltzmann entropy in order to preserve and express Shannon information.
LLMs use computation to spend physical energy in order to reduce semantic uncertainty and generate ordered meaning.
That is the bridge between the central dogma, the Krebs cycle, and artificial intelligence.
The central dogma is the code becoming form.
The Krebs cycle is the energy becoming work.
The LLM is the weights becoming meaning.
And all three point toward the same grand principle:
Information does not act by itself.
It must be expressed through an energy-driven cycle.
Wherever stored pattern meets powered transformation, something like function begins.
Leave a Reply